Age prediction in Atlantic Sturgeon
Methylation-based age prediction in Atlantic Sturgeon
 Kara Jones
Kara Jones Atlantic Sturgeon are long-lived, late-maturing fish that are endangered across most of their range. Populations face ongoing threats from ship strikes, barriers such as dams that limit access to spawning habitat, and incidental capture as bycatch in commercial fisheries. Effective conservatihttps://doi.org/10.1101/gr.266551.120on and recovery planning depend on accurate estimates of age, which are essential for understanding growth rates, survival, and population dynamics. However, traditional aging methods that rely on hard structures (such as fin rays and spines) can be unreliable in older individuals, while other structures (such as otoliths) can only be collected post-mortem.
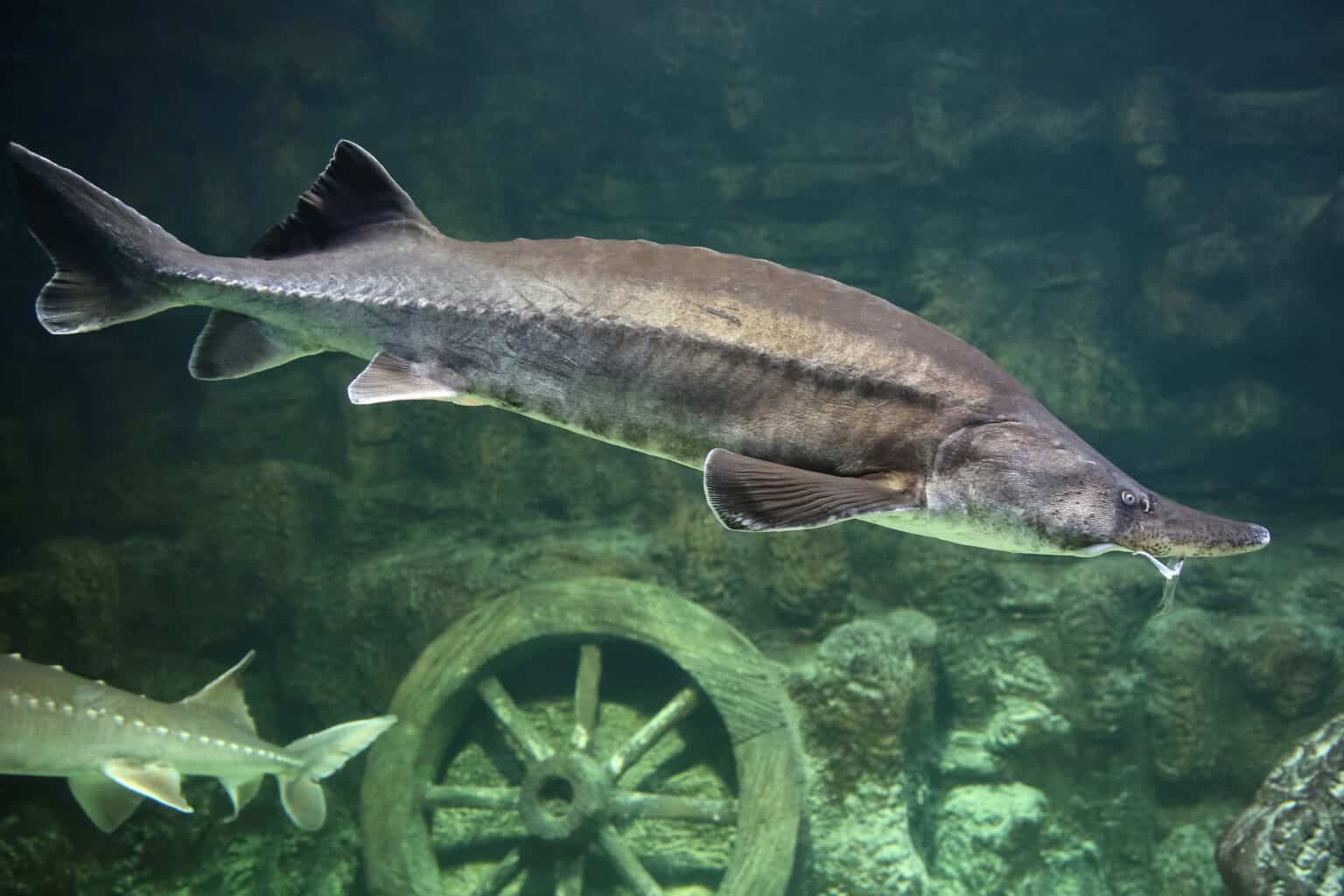 Atlantic Sturgeon (Acipenser oxyrinchus)
Atlantic Sturgeon (Acipenser oxyrinchus)
This project addresses a key limitation identified in recent stock assessments by evaluating the use of “epigenetic clocks”—a molecular approach that estimates age based on predictable changes in DNA methylation over time. Because DNA methylation can be measured from a variety of tissue types, this method offers a non-lethal, scalable, and potentially more accurate alternative to conventional techniques.
 Epigenetic Clock, from Xiao, Wang, & Kong 2019
Epigenetic Clock, from Xiao, Wang, & Kong 2019
Using Enzymatic Methyl-sequencing (EM-seq, Vaisvila et al. 2021), we will be screening tens of thousands of candidate methylation sites to develop a predictive model of age in wild Atlantic Sturgeon. By building a robust and unbiased model that performs consistently across sexes and populations, this project aims to deliver a practical, management-ready tool to improve stock assessments and support long-term conservation of this species.
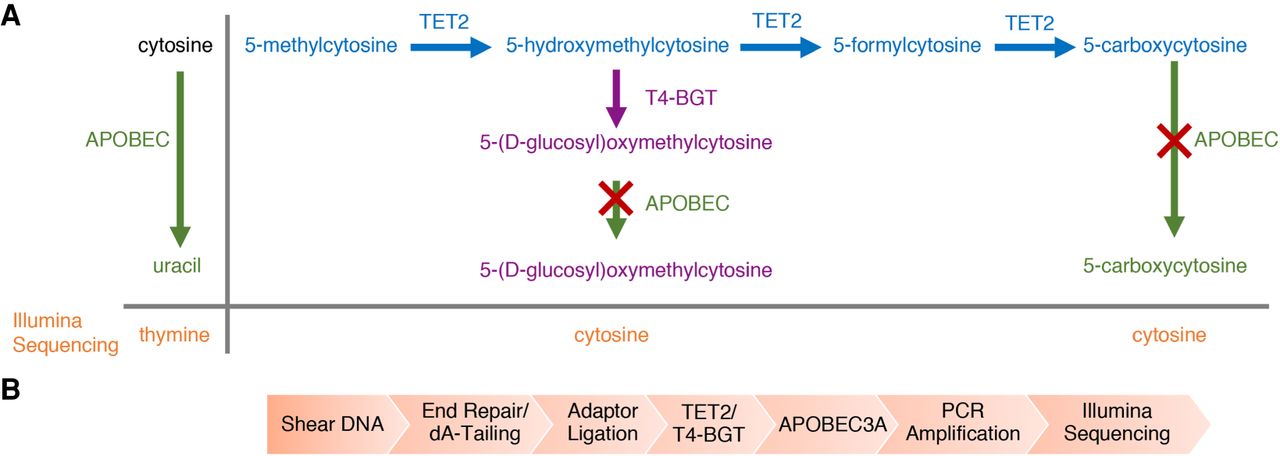 EM-Seq, from Vaisvila et al. 2021
EM-Seq, from Vaisvila et al. 2021
