Low-Coverage Whole Genome Sequencing
Trading sequencing depth for more samples and more data
 Runyang 'Nicolas' Lou
Runyang 'Nicolas' Lou Despite major declines in DNA sequencing costs, researchers still have to balance three trade-offs: how much of the genome to sequence (breadth), how deeply to sequence each sample (depth), and how many samples to include. Reduced-representation methods like RAD-seq have been widely used because they allow deep sequencing of a small portion of the genome across many individuals, enabling variant discovery and reliable genotyping.
However, RAD-seq leaves large parts of the genome unsequenced, which can cause it to miss localized signals of selection or adaptation. In many cases, whole-genome sequencing has revealed important patterns that RAD-seq failed to detect, suggesting that full genome coverage is often necessary. The downside is cost: sequencing entire genomes at high depth for many individuals remains expensive.
One cheaper alternative is pooled sequencing (Pool-seq), where DNA from multiple individuals is combined and sequenced together. This approach can accurately estimate population-level patterns but loses all individual-level information, making it harder to detect uneven contributions or hidden population structure.
Low-coverage whole-genome sequencing (lcWGS) offers a middle ground. By sequencing many individuals at low depth across the entire genome, it captures broad genomic information while retaining individual identities. This approach sacrifices confidence in individual genotypes but gains wider genome coverage and often allows larger sample sizes at a comparable cost to RAD-seq or Pool-seq.
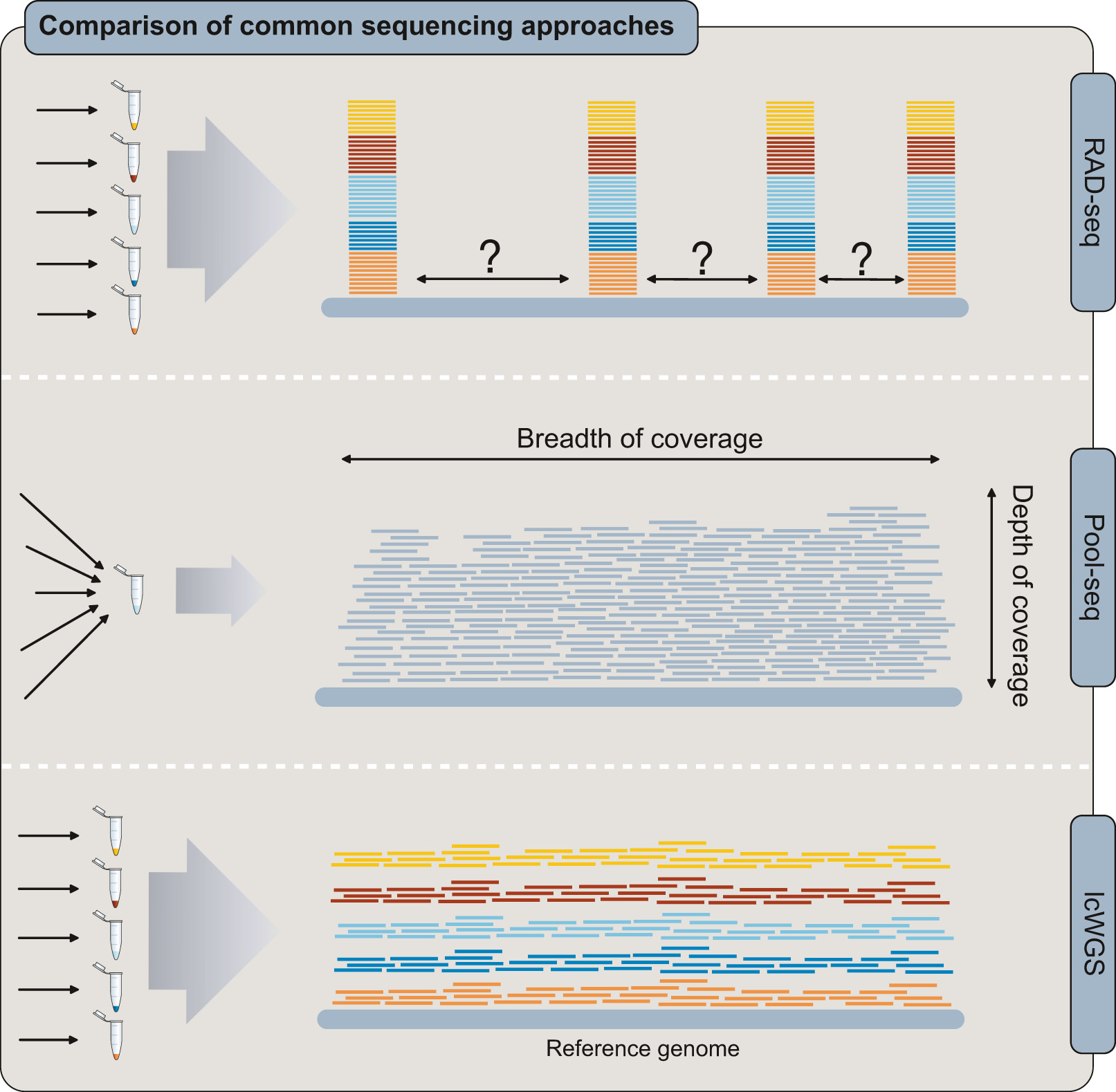 lcWGS vs other methods
lcWGS vs other methods
